Abstract
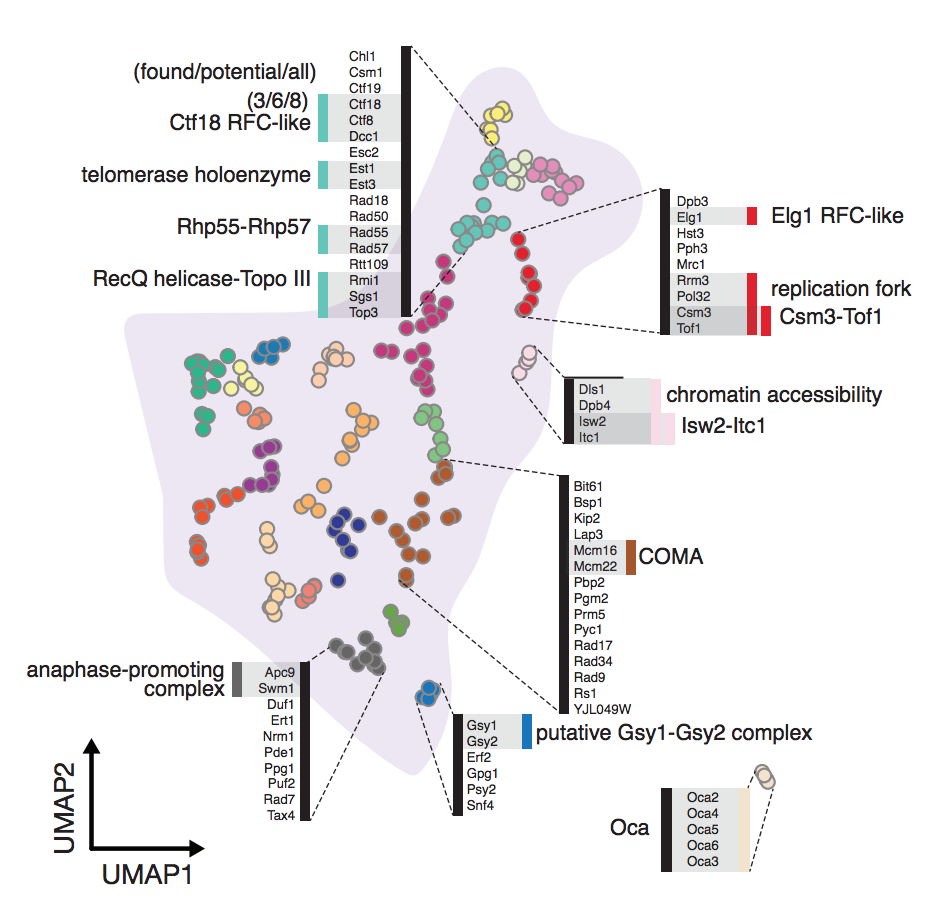
Dimensionality reduction is often used to visualize complex expression profiling data. Here, we use the Uniform Manifold Approximation and Projection (UMAP) method on published transcript profiles of 1484 single gene deletions of Saccharomyces cerevisiae. Proximity in low-dimensional UMAP space identifies groups of genes that correspond to protein complexes and pathways, and finds novel protein interactions, even within well-characterized complexes. This approach is more sensitive than previous methods and should be broadly useful as additional transcriptome datasets become available for other organisms.
* co-first authors
** corresponding authors