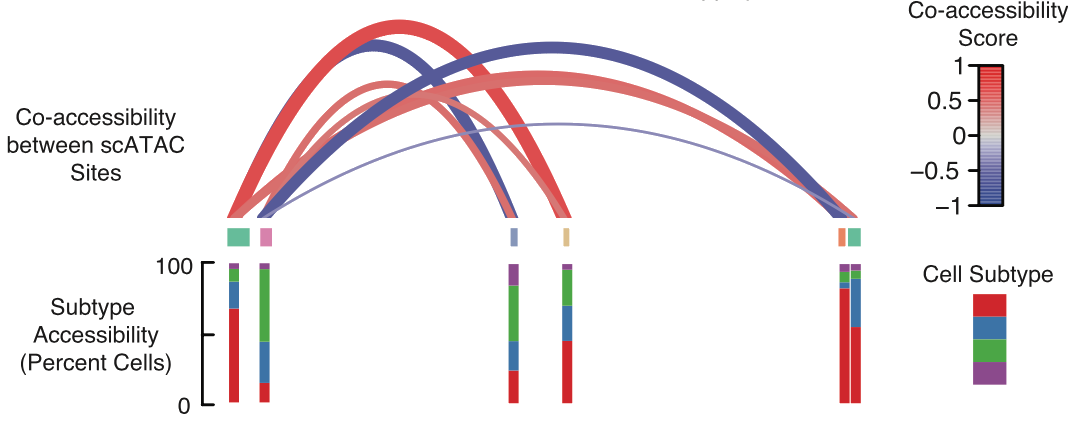
ATAC-Seq is an assay for interrogating the entire genome for accessibility to DNA binding proteins in a single experiment. In collaboration with Jay Shendure’s lab and scientists at Illumina, we recently developed sci-ATAC-seq, a single-cell ATAC-seq protocol. Our initial study, led by Darren Cusanovich, explored variation in chromatin accessibility both between and within populations of single cells. Our single-cell ATAC-Seq relies on combinatorial cellular indexing, and thus does not require the physical isolation of individual cells during library construction. The technique scales sublinearly in time and cost and can profile thousands of individual cells in a single experiment.
Analyzing single-cell ATAC-Seq experiments is challenging because the datasets tend to be large, sparse, and binary. Conventional single-cell analysis approaches such as principal component analysis are not well suited to this sort of data. We are working on new techniques to glean insights about cell state and gene regulation from single-cell ATAC-Seq experiments.